2020 Knight Campus Undergraduate Scholars Take Their Research Remote
The Knight Campus Undergraduate Scholars Program pairs promising undergraduates with research mentors — graduate students, postdocs, and faculty members — immersing them in a 12-month, comprehensive research experience in Knight Campus-affiliated labs.
For the 2020 cohort, extraordinary conditions as a result of COVID-19 moved their research from the lab to remote learning. While these young scientists are taking on independent research projects in a diverse set of fields, they are also finding creative ways to conduct their research remotely. Below are some examples of what the cohort has been working on this term.
Industrial Source Scholar
Mentor: Salil Karipott
Major: Biology
Lab: Ong Lab
Fabricating and quantifying mechanically active bone fixation devices for bone fractures to enhance regeneration
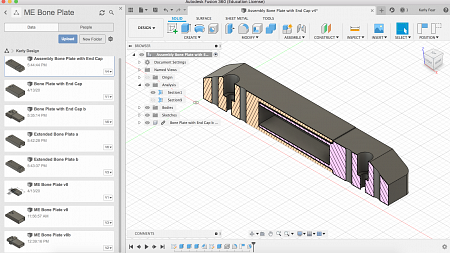
"This term, I have been using computer-aided design (CAD) software such as Fusion 360 to design the multi-component devices that my lab studies.
This is an image of a new design I am analyzing in Fusion 360. I will be designing it for a bone fixation device that I have been 3D printing, fabricating and assessing during my time as a Knight Campus Undergraduate Scholar."
Mentor: Cori Cahoon
Major: Biology
Lab: Libuda Lab
Developing biomarkers of chromosome structural defects during live sperm and egg development
"I research the biological processes of meiosis and reproduction in the Libuda Lab. Specifically, the differences in the development of sperm versus eggs.
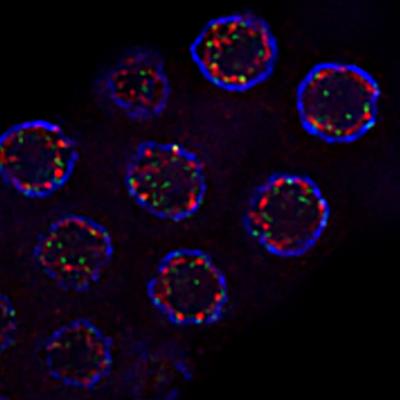
This is a photo of heat-shocked C. elegan worms that shows nuclei within the germline. The green spots mark DNA damage and allow me to quantify the amount of damage when the sample is subjected to elevated temperatures. The blue circles are staining DNA, and the red spots indicate stain Lamin, a nuclear membrane protein.
I am interested in studying why exposure to heat stress causes infertility in males. Images like these can be compared to those without heat stress conditions to understand why we see higher rates of DNA damage in sperm development than in egg development under similar conditions.
After graduation, I hope to attend medical school and become an Orthopedic Surgeon focusing on sports medicine."
Guldberg Scholar
Mentor: Deepika Sundarraman
Major: Biology
Lab: Parthasarathy Lab
Investigating the spatial organization and physical structure of gut microbes
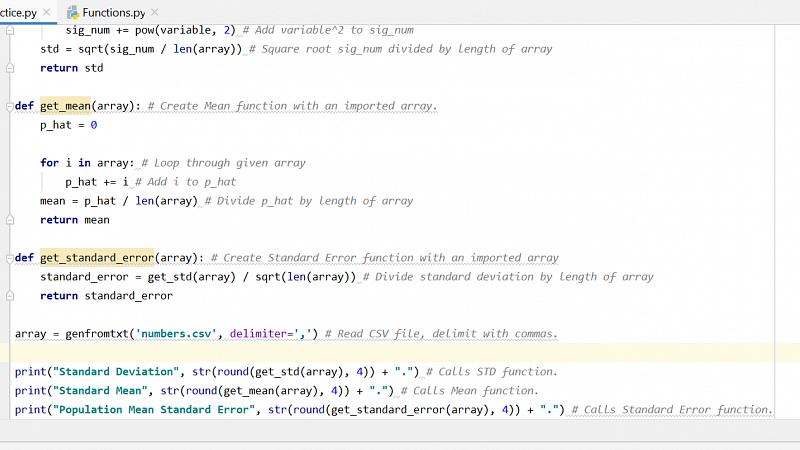
"I am learning how to program in Python to analyze the bacterial abundance data I collected at the beginning of the term.
This image shows the code I am learning to write that calculates the mean, standard deviation, and standard error of a given data set."
Petrone Scholar
Mentor: Calin Plesa
Major: Biology
Lab: Plesa Kab
Repurposing the minimal replication system of a bacteriophage as a tool for bioengineering
"Calin Plesa is teaching me new and valuable skills that are relevant to the future I want to pursue in Biology. During my experience as a Knight Campus Undergraduate Scholar in his lab, I've been learning the basics of bioinformatics. Last term, this involved designing the replication construct and cargo DNA. This term, it has involved learning basic programming and analysis of various aspects of the PRD1 replication proteins to compare them to other organisms.
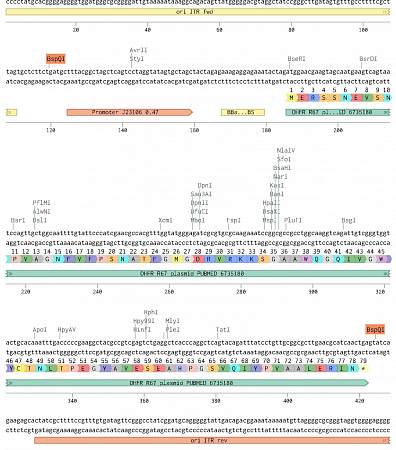
This image is an example of a linear replication construct containing the origins of replication for the bacteriophage PRD1's genome, as well as an antibiotic resistance marker (DHFR).
The bioinformatics skills I've learned through this project have helped me prepare for a career in research, and also spiked my interest in bioengineering. Professor Plesa's research goals are inspiring, and I feel fortunate to have the opportunity to work with him through the undergraduate scholars program.
After graduation, I hope to pursue a Ph.D. program in developmental biology, and eventually conduct research and teach at a research university."
Clark Honors College Scholar
Mentor: Cathy Pohl Robinson
Major: Biology with a focus on human physiology
Lab: Guillemin Lab
Constructing a SpdE-NarQ fusion protein as a biosensor for amino acids
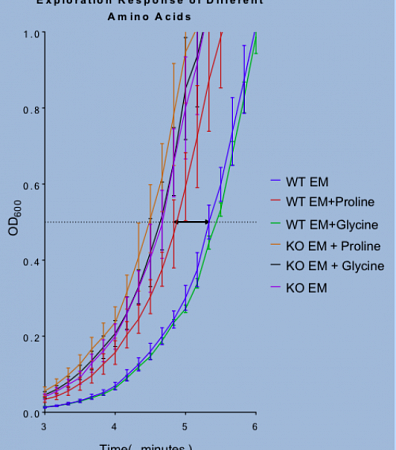
"This image shows some data I have compiled from one of my experiments as a Knight Campus Undergraduate Scholar. The data show the relationship between SpdE-dependent Aeromonas motility and environmental amino acids.
We developed a new technique we call the “exploration assay” to measure differences in motility between strains or conditions. Using the exploration assay, we compared the motility of wild type and spdE knockout strains in different amino acid environments. From our results, we found that the wild type strain is more motile in the presence of these amino acids. However, even in the absence of an amino acid signal, the spdE knockout is more motile than the wild type strains.
My focus as a Knight Campus Undergraduate Scholar is to create a model for how SpdE regulates motility in response to amino acids to offers novel insights into Aeromonas biology and the mechanisms of host colonization through my thesis project."
Mentor: Kelsey Schultz
Major: Human Physiology
Lab: Swann Lab
Analyzing the relationship between impulsivity and motor response variability
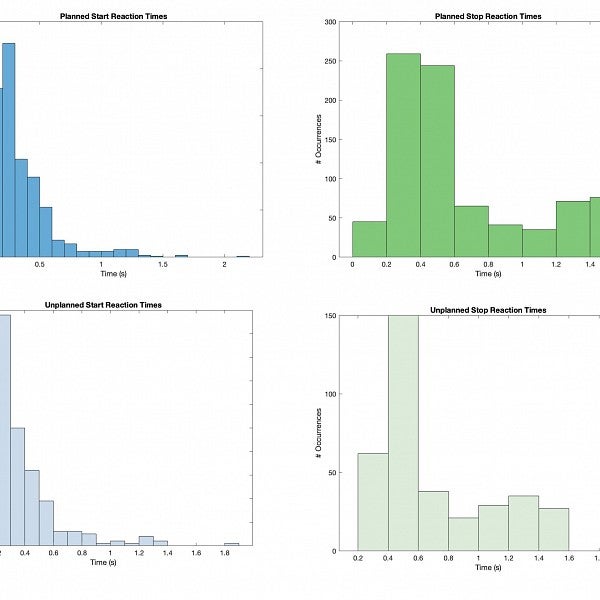
"This term, I have been preparing raw data for statistical analysis and visual plotting. This image is a histogram showing stop-signal reaction time occurrences. Stop-signal reaction times are neurocognitive tasks that provide a sensitive measure of the time taken by the brain to inhibit or suppress motor responses.
I will be using the stop-signal reaction time data and other data yet to be collected to determine the variance of planned versus unplanned signal reaction times and go signal versus stop-signal reaction times."
Mentor: Christine McDevitt
Major: Math and Computer Science
Lab: DeRose Lab
Investigating platinum induced nucleolar stress through the development of a click-capable oxaliplatin mimica
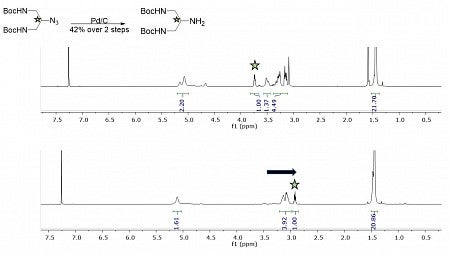
"My research in the DeRose Lab mainly entails chemical synthesis, where I run experiments to produce particular molecules. This image uses nuclear magnetic resonance (NMR) data to depict an experiment I did last term. The star indicates a specific proton in the starting material and resulting product at the location shown in the upper left. The successful trial is demonstrated by the marked shift in the proton.
This term, I have been working to develop skills such as reading NMRs and reviewing scientific papers. I also have begun to expand my conceptual knowledge of the research I hope to begin over the summer."
Clark Honors College Scholar
Mentor: Erik Toraason
Major: Biology
Lab: Libuda Lab
Defining mechanisms regulating accurate meiotic DNA repair
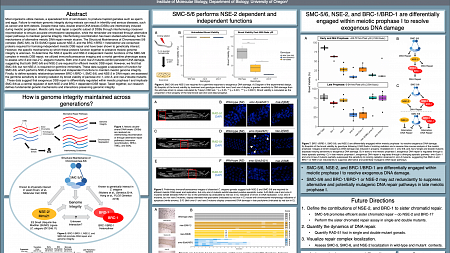
"I created this poster (click on it to enlrage) to present the research I am conducting in the Libuda Lab. I am elucidating the mechanisms that regulate DNA damage repair pathway choices during meiosis, which is crucial for preserving genomic integrity across generations. My work focuses specifically on defining the genetic interactions between three conserved proteins implicated in DNA repair: the Structural Maintenance of Chromosomes 5/6 complex, the E3 SUMO ligase NSE-2, and the BRC-1/BRD-1 heterodimer.
Over the last two weeks, I have been attending the virtual 2020 Allied Genetics Conference (TAGC), and presenting my research with the international genetics research community through a virtual poster presentation."
Mentor: Raina Krivina
Major: Chemistry
Lab: Boettcher Lab
Atomic control of the OER catalyst architecture for tuning stability and activity of the catalyst and unraveling the mechanism of OER
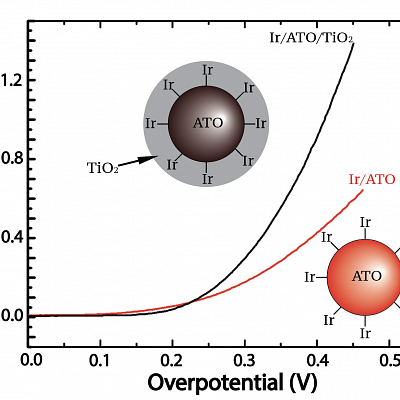
"This is an image of a cyclic voltammetry experiment with two different catalysts we've made for the oxygen evolution reaction (OER), which is rate-determining in water splitting. For the red trace, we used an iridium precursor to deposit an ultra-thin layer of iridium (the state-of-the-art catalyst or OER) on antimony doped tin oxide (ATO) as a conductive support. The black trace is the same catalyst with an additional layer of TiO2. These novel catalysts decrease the loading of iridium needed to run this reaction efficiently, making water splitting more economically feasible for clean energy storage. This plot is showing that the TiO2 enhances the performance of the catalyst, which can be seen by the curve going up at a lower overpotential and reaching higher activity levels.
Over the past couple of weeks, I've been going through the literature in my field to examine ways to move this project forward. I've also played a role in editing and drafting figures for a perspective paper our lab is planning to publish."